Abstract:
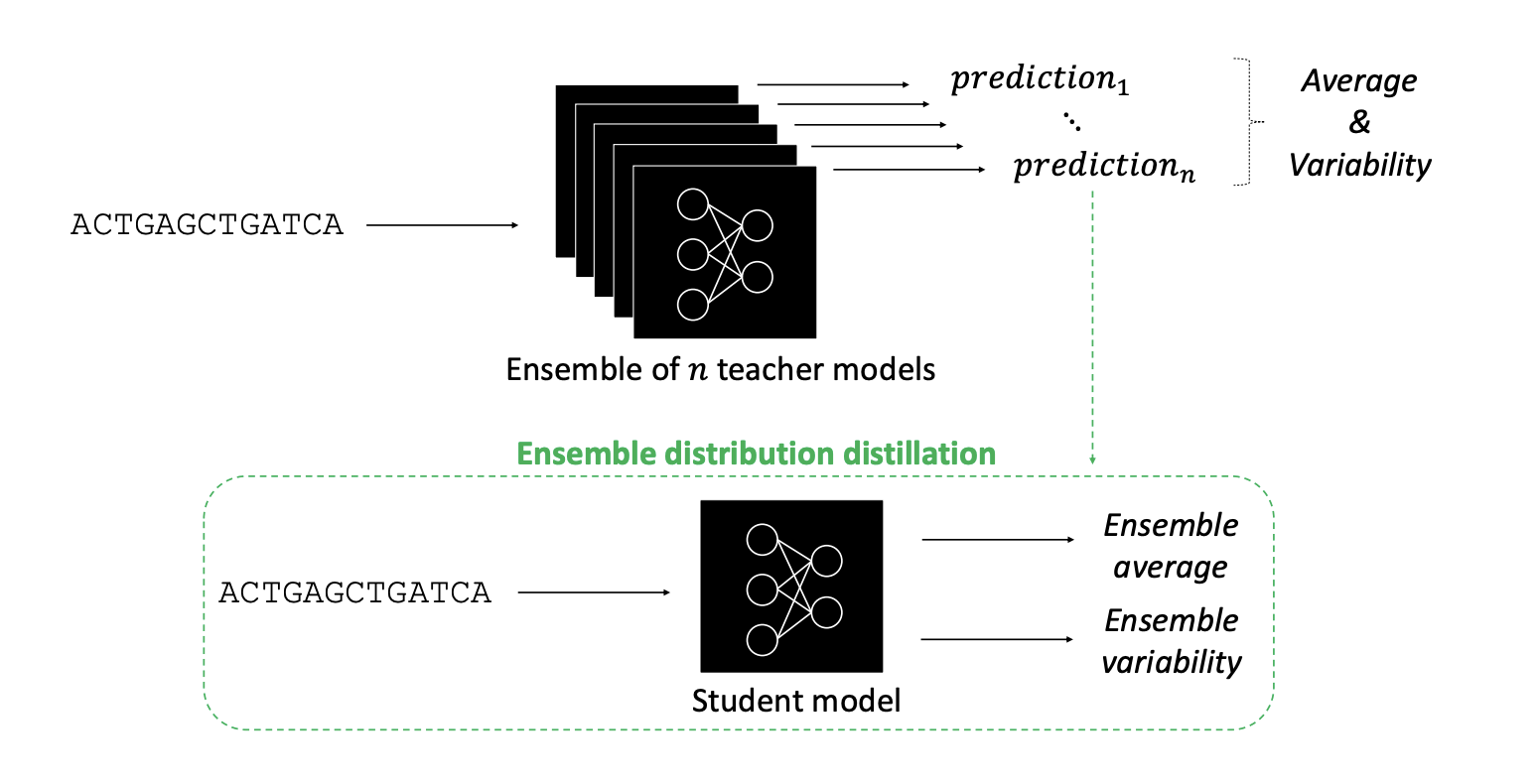
Deep neural networks (DNNs) have advanced predictive modeling for regulatory genomics, but challenges remain in ensuring the reliability of their predictions and understanding the key factors behind their decision making. Here we introduce DEGU (Distilling Ensembles for Genomic Uncertainty-aware models), a method that integrates ensemble learning and knowledge distillation to improve the robustness and explainability of DNN predictions. DEGU distills the predictions of an ensemble of DNNs into a single model, capturing both the average of the ensemble’s predictions and the variability across them, with the latter representing epistemic (or model-based) uncertainty. DEGU also includes an optional auxiliary task to estimate aleatoric, or data-based, uncertainty by modeling variability across experimental replicates. By applying DEGU across various functional genomic prediction tasks, we demonstrate that DEGU-trained models inherit the performance benefits of ensembles in a single model, with improved generalization to out-of-distribution sequences and more consistent explanations of cis-regulatory mechanisms through attribution analysis. Moreover,DEGU-trained models provide calibrated uncertainty estimates, with conformal prediction offering coverage guarantees under minimal assumptions. Overall, DEGU paves the way for robust and trustworthy applications of deep learning in genomics research.
]]>